Cytological samples obtained by endobronchial ultrasound (EBUS) are capital for diagnosis, staging and molecular profile in non-small cell lung carcinoma (NSCLC).
ObjectiveTo assess the success rate of complete, partial and individual of molecular analysis in samples obtained by EBUS-guided transbronchial needle aspiration (TBNA) and/or by oesophageal ultrasound-guided fine needle aspiration with an echobronchoscope (EUS-B-FNA) in patients with NSCLC.
MethodsProspective study including 90 patients with non-squamous NSCLC, or non-smoking squamous. Cytological samples were classified into two groups. Group 1: PEN membrane slide and/or cell blocks for the determination of mutations of EGFR, KRAS, ERBB2 and BRAF. Group 2: silane coated slides or cell blocks for rearrangements of ALK, ROS1 and MET amplification.
ResultsThe success rate was 78.6% for 4 molecular alterations (EGFR, KRAS, ALK and ROS1), and 44% for 7 determinations. The individual success rate for EGFR was 97%, KRAS 96.3%, ALK 85%, ROS1 82.3%, ERBB2 71.4%, BRAF 67.7% and MET 81.1%. There were no significant differences (p=0.489) in the number of molecular analyses (1–3 vs. 4) in group 1, depending on the types of samples (cell block vs. PEN membrane slide vs. cell block and PEN membrane slide).
ConclusionsIn patients with NSCLC, the cytological material obtained by ultrasound-guided needle aspiration is sufficient for individual and partial molecular analysis in the vast majority of cases. Membrane slides such as cell blocks are valid samples for molecular analysis.
Lung cancer is the most frequent malignant neoplasm and the main cause of cancer mortality.1 NSCLC accounts for 85% of all lung cancers, with adenocarcinoma as the most prevalent histological subtype.1,2 Currently, the identification of different oncogenic alterations leads to oncospecific treatment which improves the overall response rate and progression-free survival in this group of patients.3,4
The diagnostic and/or staging process includes different minimally invasive and invasive techniques to obtain cytological and/or histological material, which provides differentiation of the histological subtype and identification of molecular markers according to treatment guidelines.5,6 EBUS-TBNA is a cost-effective tool for obtaining cytological samples safely, less invasively, and with a lower complication rate, when compared to surgical techniques.7,8 Several studies have shown that the performance of isolated molecular determinations is feasible in 70–90% of samples obtained by EBUS-TBNA.9,10 Nevertheless, for newer onco-targeted treatments, a greater amount of cyto/histological material is required in order to analyse a greater number of possible molecular targets or resistances.
The main objective of our study was to assess the success rate in the determination of molecular alterations (EGFR – epidermal growth factor receptor, KRAS – kirsten rat sarcoma viral oncogene homolog, ALK – anaplastic lymphoma receptor tyrosine kinase, ROS1 – proto-oncogene tyrosine-protein kinase ROS, ERBB2 – erb-b2 receptor tyrosine kinase 2, BRAF – v-Raf murine sarcoma oncogene homolog B1 and MET – tyrosine-protein kinase Met) in a complete, partial and individual way, from samples obtained by EBUS-TBNA and/or EUS-B-FNA in patients with NSCLC. The secondary objectives of this study were to study its prevalence in our population, and to compare the usefulness of different cytological samples for mutation analysis.
MethodsPatientsThis is an observational prospective study conducted in a tertiary hospital in patients with suspected or known NSCLC undergoing EBUS-TBNA and/or EUS-B-FNA for diagnosis and/or staging study, from January 2013 to December 2016. The inclusion criteria were patients with advanced stages (IIIB, IV), or stage IIIA non-tributary of surgical treatment, and cases with suspected recurrence or disease progression, in the absence of an active treatment.
Current smokers with squamous carcinoma were excluded. Our study was approved by the Ethics Committee of our hospital (PI-14-073), and the patients signed informed consent forms.
ProceduresTBNA or FNA were performed under local anaesthesia with lidocaine, and with moderate sedation with midazolam, propofol and/or remifentanil. The convex probe EBUS (UC 180F, Olympus Optical Co Ltd., Tokyo, Japan) was used, and sampling was performed with the 22-gauge cytology needle (NA2015X-4022; Olympus Optical Co, Tokyo, Japan). Mediastinal and/or hilar nodes with a diameter of the minor axis ≥5mm and central masses were punctured. The bronchoscopist proceeded from nodes in regions corresponding to N3 disease to regions of N1 disease.8 We performed 1–7 punctures per node and/or mass with a maximum of 15 revolutions per puncture, and negative pressure was used (−20cmH2O) in most cases.
Preparation of cytological samplesThe aspirates obtained were placed on slides, fixed with 96% ethanol and stained with haematoxylin. Rapid on-site evaluation (ROSE) was performed by a cytopathologist. Aspirates were considered satisfactory with the following criteria: (1) Benign: 40 lymphocytes per high-power field in cellular areas of the smear and/or clusters of pigmented macrophages without evidence of malignant cells or (2) Metastatic: presence of malignant cells. Samples were considered inadequate if they showed only bronchial/oesophageal cells, erythrocytes or necrotic tissue.
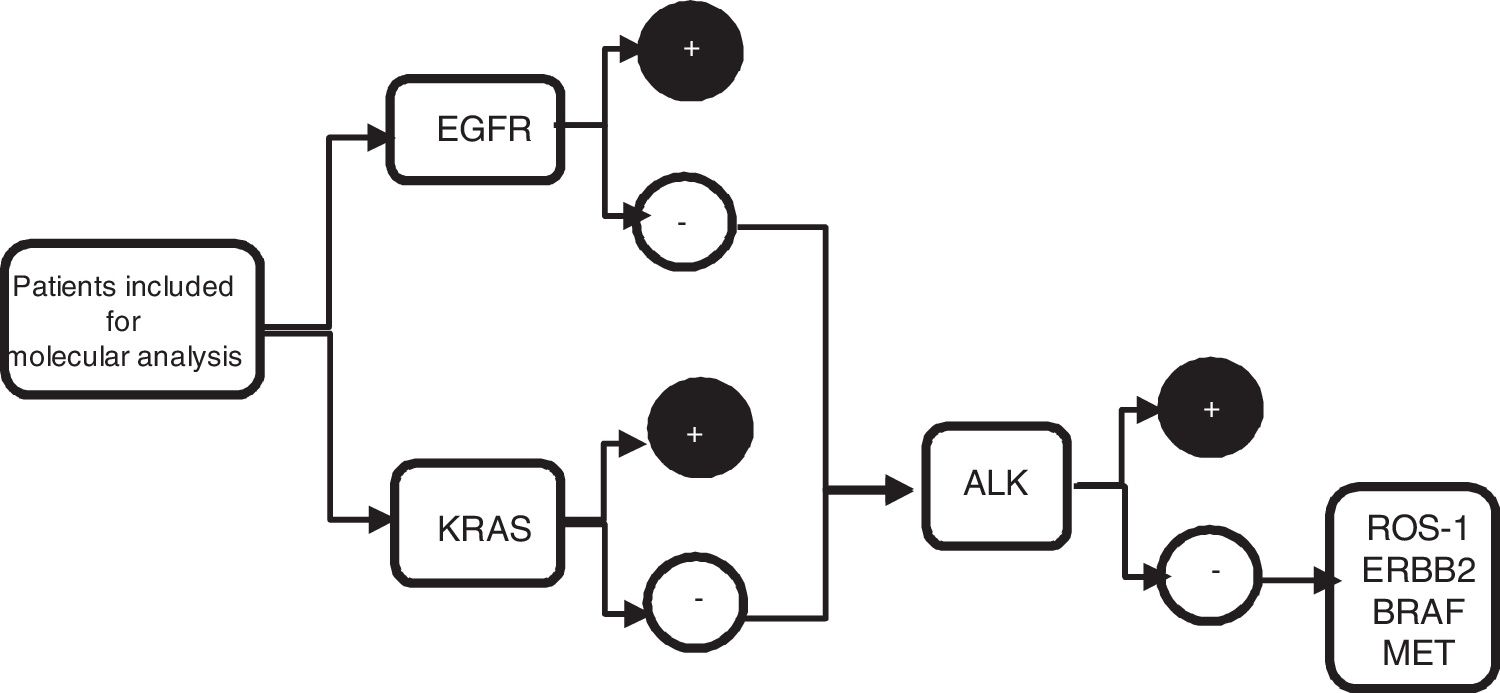
From the metastatic aspirate, two cell blocks±PEN (polyethylene naphthalate) membrane slide (Carl Zeiss, membrane slide NF 1.0 PEN, Germany) were prepared, and if it was not feasible (even with procoagulants), 4 extensions were performed on silane coated slides, and 1 PEN membrane slide. The cell blocks were prepared by air-drying the slides to clot, fixed in 10% formalin, and subsequently analysed in laboratory. Blocks were included in paraffin and sectioned (4μm thickness, 8–10 slides), two for with haematoxylin–eosin staining, two slides for immunohistochemistry staining (IHC) such as thyroid transcription factor 1 (TTF-1) and/or p40, the remaining slides were later used for sequential molecular analysis (Fig. 1).
Algorithm of sequential molecular analysis. EGFR: epidermal growth factor receptor; KRAS: kirsten rat sarcoma viral oncogene homolog; ALK: anaplastic lymphoma receptor tyrosine kinase; ROS1: protooncogene tyrosine protein kinase ROS; ERBB2: erb-b2 receptor tyrosine kinase 2; BRAF: v-Raf murine sarcoma oncogene homolog B1; MET: tyrosine-protein kinase Met.
The samples obtained were divided into two groups. Group 1: PEN membrane slide and/or cell blocks and were used to perform PCR for molecular analysis of EGFR, KRAS, BRAF and ERBB2. Group 2: silane coated slide or cell blocks and were used to perform FISH techniques for ALK, ROS1, and MET studies.
Laboratory methodsDetection of EGFR, KRAS, BRAF, ERBB2 genes mutationsThe polymerase chain reaction (PCR) technique was used from cell blocks or membrane slides, where tumour cell extraction was performed using laser microdissection techniques (Carl Zeiss MicroImaging GmbH, München, Germany). DNA was extracted with phenol–chloroform–isoamyl alcohol. If tumour cells were <200, a lysis buffer with proteinase K compatible with the PCR buffer was used (Ecogen, Barcelona, Spain). For the determination of deletions of exon 19 of the EGFR, the GeneScan technique (Applera, Norwalk, CT, USA) was used, while for the detection of the L858R mutation of exon 21 and T790M of exon 20 the Taqman technique was used (Applied Biosystems).
Genomic KRAS codons 12 and 13 mutations were assessed by the Sanger technique. Taqman technique was used for V600 mutations in BRAF and exon 20 in ERBB2.
Detection of ALK, ROS1 genes rearrangements and MET gene amplificationFluorescence in situ hybridisation (FISH) was used with the following probes: the Vysis ALK Break-Apart (Abbott Molecular, Inc., Des Plaines, IL, USA) for ALK, ZytoLight® SPEC ROS1 Dual Color Break Apart (ZytoVision, Bremerhaven, Germany) for ROS1, and ZytoLight® MET/CEP 7 Dual Color (ZytoVision, Bremerhaven, Germany) for MET. After dewaxing, the slides were processed with the Histology FISH Accessory kit (Dako, Denmark) and hybridisation titration was carried out with an Olympus BX51 fluorescence microscope. A minimum of 50 tumour cells was necessary. For ALK and ROS1 genes rearrangements, samples were considered positive when ≥15% of the cells showed a positive hybridisation pattern. For the MET gen, the criteria described by Noonan et al.11 was used according to a ratio of MET/centromere ≥1.8.
Statistical analysisCategorical variables were expressed in relative and absolute frequencies; continuous variables in mean and standard deviation when they presented a normal distribution, or as median and interquartile range when they did not present a normal distribution.
The success rate for the complete or individual molecular analysis in the samples obtained by EBUS-TBNA or EUS-B-FNA was calculated. The relationships between categorical variables were analysed using the chi-square test or Fisher's exact test. Values of p<0.05 were considered statistically significant. Data were analysed using IBM SPSS software version 24.0 (SPSS Inc., Chicago, IL, USA).
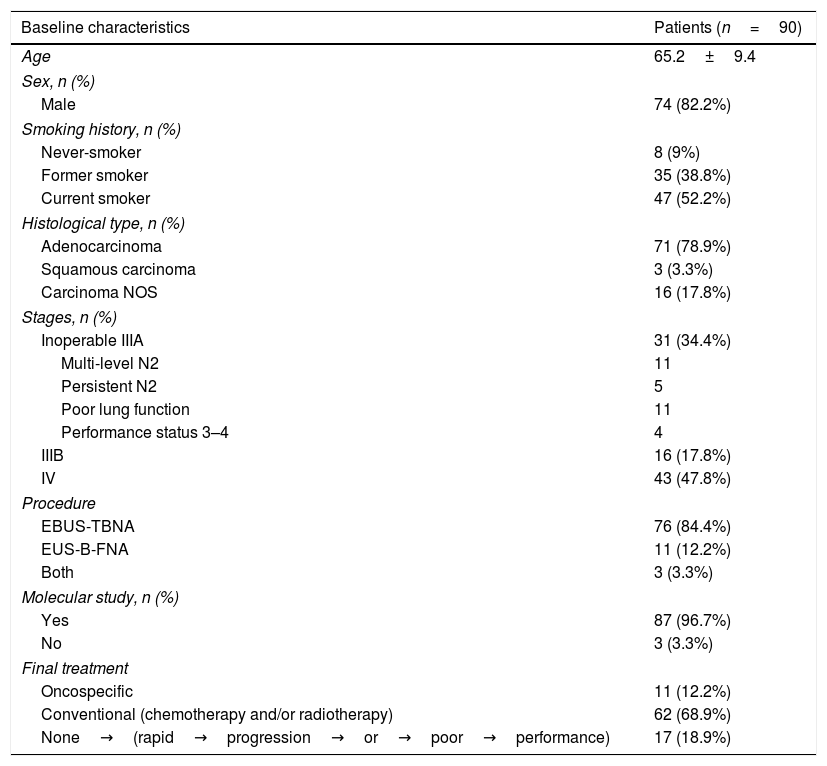
ResultsPatient characteristicsNinety patients were included in the study. In 54 cases (60%), EBUS/EUS-B was the first diagnostic procedure performed. Seventy-one patients (78.9%) had a diagnosis of adenocarcinoma. Baseline characteristics are shown in Table 1.
General characteristics of patients.
| Baseline characteristics | Patients (n=90) |
|---|---|
| Age | 65.2±9.4 |
| Sex, n (%) | |
| Male | 74 (82.2%) |
| Smoking history, n (%) | |
| Never-smoker | 8 (9%) |
| Former smoker | 35 (38.8%) |
| Current smoker | 47 (52.2%) |
| Histological type, n (%) | |
| Adenocarcinoma | 71 (78.9%) |
| Squamous carcinoma | 3 (3.3%) |
| Carcinoma NOS | 16 (17.8%) |
| Stages, n (%) | |
| Inoperable IIIA | 31 (34.4%) |
| Multi-level N2 | 11 |
| Persistent N2 | 5 |
| Poor lung function | 11 |
| Performance status 3–4 | 4 |
| IIIB | 16 (17.8%) |
| IV | 43 (47.8%) |
| Procedure | |
| EBUS-TBNA | 76 (84.4%) |
| EUS-B-FNA | 11 (12.2%) |
| Both | 3 (3.3%) |
| Molecular study, n (%) | |
| Yes | 87 (96.7%) |
| No | 3 (3.3%) |
| Final treatment | |
| Oncospecific | 11 (12.2%) |
| Conventional (chemotherapy and/or radiotherapy) | 62 (68.9%) |
| None→(rapid→progression→or→poor→performance) | 17 (18.9%) |
Carcinoma NOS (not otherwise specified) type; EBUS-TBNA (endobronchial ultrasound guided biopsy); EUS-B-FNA (oesophageal ultrasound-guided fine needle aspiration with an echobronchoscope).
Of the 12 pulmonary lesions (24–63mm of short-axis diameter) and 329 hilar and/or mediastinal lymphadenopathies punctured, 114 (30.6%) were malignant. The most frequently punctured metastatic lymph nodes stations were 7 (33), 4R (30), 4L (12) and 10/11/12 (11). The mean short-axis diameter of the lymph node was 13.3±6.8mm and the average number of punctures per node was 3 (range 1–7).
Histological subclassificationIn 35 cases (39%), the final diagnosis was obtained by morphological assessment, while IHC (TTF-1, p40) was used in 53 (58.9%). A definitive subtyping (morphology and/or IHC) was achieved in 82.2% vs. 17.8% of carcinoma NOS (not otherwise specified). In this subgroup of 14 cases, both morphological assessment and IHC were inconclusive, while in 2 cases IHC was not performed because the material for the molecular analysis was prioritised.
Molecular analysisIn 96.7% cases it was feasible to start the sequential molecular analysis. The samples obtained were insufficient in the remaining 3 cases: one corresponded to re-staging (difficulty obtaining viable material post neoadjuvant inflammatory changes), and silane coated slides and PEN membrane slides did not contain enough tumour cells for molecular detection in the other two. In the first case, the patient underwent palliative treatment due to deterioration of general condition, while in the other two cases a second procedure was performed [puncture of the primary mass by CT-guided percutaneous transthoracic needle biopsy (CT-guided PTNB) and by EBUS].
IHC was performed in both sufficient and insufficient samples for complete molecular analysis (65.5% vs. 60%, p=0.115).
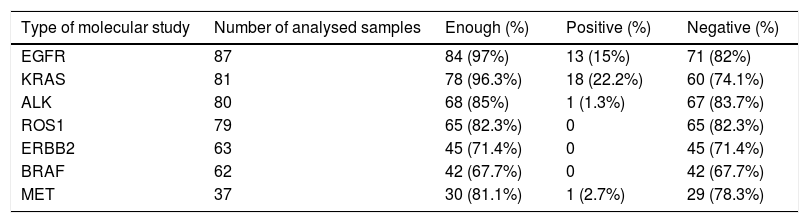
A partial molecular analysis (EGFR, KRAS, ALK and ROS1) was performed in 78.6% of patients, while complete analysis was carried out in 44% of cases. Table 2 details the individual results for each molecular detection.
Individual molecular analysis.
| Type of molecular study | Number of analysed samples | Enough (%) | Positive (%) | Negative (%) |
|---|---|---|---|---|
| EGFR | 87 | 84 (97%) | 13 (15%) | 71 (82%) |
| KRAS | 81 | 78 (96.3%) | 18 (22.2%) | 60 (74.1%) |
| ALK | 80 | 68 (85%) | 1 (1.3%) | 67 (83.7%) |
| ROS1 | 79 | 65 (82.3%) | 0 | 65 (82.3%) |
| ERBB2 | 63 | 45 (71.4%) | 0 | 45 (71.4%) |
| BRAF | 62 | 42 (67.7%) | 0 | 42 (67.7%) |
| MET | 37 | 30 (81.1%) | 1 (2.7%) | 29 (78.3%) |
EGFR: epidermal growth factor receptor; KRAS: kirsten rat sarcoma viral oncogene homolog; ALK: anaplastic lymphoma receptor tyrosine kinase; ROS1: protooncogene tyrosine protein kinase ROS; ERBB2: erb-b2 receptor tyrosine kinase 2; BRAF: v-Raf murine sarcoma oncogene homolog B1; MET: tyrosine-protein kinase Met.
Mutations in the EGFR genes were detected in thirteen cases (15%), 12 of them (92.3%) belonging to the adenocarcinoma subtype. Mean age was 67.8±10 years, 38% were women, and 5 cases corresponded to never-smokers. The most frequent mutation was the deletion of exon 19 (54%).
Molecular analysis by groupIn group 1, it was feasible to carry out 4 determinations in 38 (44%), while only 1–3 mutations could be studied in 49 (56%). There were no significant differences in the number of molecular determinations (1–3 vs. 4) (p=0.489) when separated by sampling subgroups between: (1) cell block, (2) PEN membrane slide, and (3) cell block and PEN membrane slide (Table 3).
In group 2, it was feasible (70/81) to carry out 3 determinations (ALK and/or ROS1 and/or MET) in 86%. Of these, complete analysis was performed in 28 cases (34.6%), and only 2 in 38 cases (47%).
DiscussionThe results of our study showed that the cytological samples obtained by EBUS-TBNA or EUS-B-FNA, mainly cell blocks and PEN membrane slides, has had a high success rate for sequential molecular analysis, both partially and individually. The College of American Pathologists (CAP), the International Association for the study of Lung Cancer (IASLC) and the Association for Molecular Pathology (AMP) jointly issued guidelines for routine testing of biomarkers in lung cancer and recommended performing the EGFR, ALK and ROS1 genes rearrangements as a first step, and KRAS only if the studies of the first step are negative.6 In our series, the partial success rate with these 4 biomarkers was around 80%. However, the individual success rate decreased with the sequential analysis, so that EGFR detection was 96.6% while for BRAF it was 67.7%. These results could be explained by the loss of tumour material during the cell block cuts, scraping tumour cell from PEN membrane slide by manual or laser microdissection, or by the delay in processing fresh slides.12
The usefulness of cytological samples obtained by EBUS/EUS-B for molecular analysis has been assessed in different studies, with 2 or 3 molecular alterations studied in most of them.9,10,13–16 In our series, by applying a protocol for obtaining cytological material, effectiveness was optimal (96.7%) to start sequential molecular analysis, and it was up to 44% favourable for 7 biomarkers. Jurado et al.13 observed that in 82% (42/56) of cases, enough cytological material was obtained for molecular testing (EGFR, ALK and KRAS), while in our study there were 4 detections (EGFR, KRAS, ALK and ROS1) in 78.6% of cases, which was similar but with an additional biomarker in our series. Folch et al.16 compared the detection of 3 biomarkers (EGFR, KRAS and ALK) in cytological samples (cell blocks) obtained by EBUS, with samples obtained by other techniques (surgical biopsy of mediastinal and hilar nodes, bronchial biopsies, CT-guided PTNB of lung lesions) and managed to show a success rate above 90%, confirming that the EBUS is a suitable tool with which to initiate a genotypic study of NSCLC.
The presence of ROSE was a notable advantage in the optimisation and handling of the samples, as it increases performance in obtaining tumour material for IHC and molecular testing.17–19 Trisolini et al.20 demonstrated that the presence of an anatomopathologist can prevent the performance of additional procedures for genotyping in 1 in 10 cases and, although there was no statistical relevance, clinical relevance was observed. In our institution, the good cooperation between cytopathologist and bronchoscopist facilitated the preparation of the samples with adequate tumour material, and in some cases, allowed us to modify the puncture zone in the metastatic lymph node (e.g. presence of necrosis zones), change the lymph node station, or the number of punctures. In our study, ROSE allowed us to perform an average of 3 punctures to obtain sufficient material, below the 4 punctures recommended by the guidelines for obtaining material for molecular study,21 yet our success rate for partial study was close to 80%.
When evaluating by type of cytological sample (cell block±PEN membrane slide) for the number of mutations (1–4) we did not observe differences between these samples. PEN membrane slides are an additional tool that contains DNA for molecular subtyping in NSCLC.
The success rate for the detection of EGFR in cytological samples obtained by EBUS varies between 72.2% and 98.7%.9,10 In our sample, it was 97%, higher than the 90% published by Navani et al.15 In our study, mutation in EGFR genes was detected in 15%, similar to that reported in histological samples from other Spanish studies (11.6–16.6%).3,22
The success rate for the detection of KRAS was 96.3%, higher than the results reported in the studies by Kang and Jurado, 90.2% and 75% respectively13,23. Mutations in the KRAS genes were observed in 20%, similar to that reported (23.6%) in different cytological series.24 In line with previous research, in our experience the presence of EGFR and KRAS mutations was exclusive.24
The determination of ALK-rearrangements was possible in 85% of samples, similar to the experience of Jurado et al.,13 which was 91% (39/41) with a frequency of positive cases (by FISH) of 6.4% (7/109),14 while in our group only 1 case was positive.
Our article shows that it is feasible to perform a greater number of detections in cytological samples such as cell blocks and membrane PEN slides, following a sequential algorithm and with the help of a cytopathologist. The limitations of our study includes not having all the determinations available in all cases (our assistance protocol did not perform additional analysis in cases where a molecular alteration was already detected) and it was based on data from a single centre with a small sample size, which limits the generalizability of our results.
Immunotherapy with checkpoint inhibitors was recently approved to treat NSCLC. Wang et al.25 demonstrated 87% feasibility to determine the expression of PD-L1 (programmed death ligand 1) in samples obtained by EBUS-TBNA with a good correlation between the cytological and surgical samples. Likewise, the incorporation of second-generation sequencing (NGS) allowed the analysis of numerous genetic alterations simultaneously, with the use of small amounts of DNA (nanograms). However, its usefulness in cytological samples obtained by EBUS should be validated, since it requires quality DNA and a sufficient tumour cell percentage to give adequate sequencing.26
In conclusion, in our series applying a protocol based on obtaining a minimum of 2 cell blocks and optimising the handling of the samples for multiple molecular analysis, we have shown that the cytological samples obtained by EBUS-TBNA or EUS-B-FNA are suitable in a high percentage of patients for individual and partial molecular analysis. PEN membrane slides and cell blocks have shown to be equally valid samples for the determination of mutations.
FundingThis study was funded by a research grant from the Spanish Society of Pulmonology and Thoracic Surgery (SEPAR 198/2013), Antonio Castellá Award for research projects of the Spanish Association of Respiratory Endoscopy (AEER) and PUBLIBECA of SEPAR. Additionally, it received support from the Germans Trias Talents – 2013 Aid Program, Germans Trias i Pujol Hospital and Institute.
Conflicts of interestThe authors have no conflicts of interest to declare.
To Jana Pagès Barón, cytotechnician in our centre for her invaluable collaboration to achieve the appropriate sample circuit between the Molecular Biology Laboratory of the ICO and our centre.